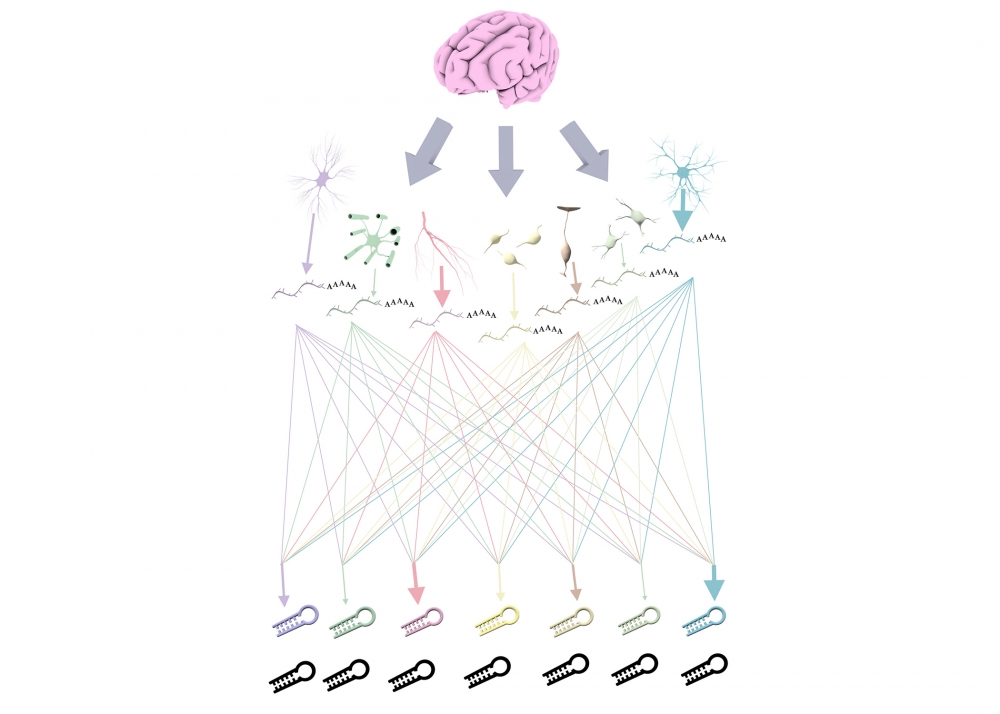
The Builder and the Regulator


One of the big challenges in neurobiology is cell classification, a problem compounded by the fact that the same cell type can look different depending on the method of analysis used to classify it — whether by cell shape, gene expression profile, electrophysiological firing pattern, or selective vulnerability to certain diseases.
“There are many individual cell types in the brain, and we don’t know how to identify them all,” said Kenneth S. Kosik, Harriman Professor of Neuroscience in the Department of Molecular, Cellular and Developmental Biology at UC Santa Barbara and co-director of the campus’s Neuroscience Research Institute. In classification paradigms that focus on which genes express themselves to define cell type, researchers have looked at messenger RNAS (mRNAs). But in a project described in the paper “Regulation of Cell Type–Specific Transcriptomes by MicroRNA Networks During Human Brain Development,” Kosik and his co-authors “brought an additional parameter — micro-RNAs (miRNAs) — to the problem of classification. Researchers had not previously looked at miRNAs at the single-cell level to get at cell-type specificity.” If DNA is the architect holding the genetic blueprint for species replication, then RNA is the on-the-ground builder, responsible sometimes for initiating, and other times for carrying out or delegating, multiple processes essential for creating fully realized living forms.
Key tasks for which messenger RNA is responsible include reading genetic information from DNA and translating it into proteins that fire up the cell-production factory. RNA also plays a role in regulating the cell-production process, an especially important function for ensuring that genes are expressed correctly and that cells are produced in the correct amounts.
In its regulatory role, mRNA has an indispensable assistant — microRNA (miRNA) — which works to prevent over-proliferation of specific types of cells. Such overproduction “can have implications in diseases that have prenatal origins, such as ASD [autism spectrum disorder],” said co-first author Mahdi Golkaram, a bioinformatics scientist at illumina who earned a Ph.D. in mechanical engineering at UC Santa Barbara in 2018.
The interactions between mRNA and miRNA are the subject of the paper. The researchers sought to understand the complex time- and context-specific interplay between the two, and how imbalances between them may relate to prenatally formed disease in the human brain.
“Better understanding how mRNAs and miRNAs work together, and how that looks in normal cell development and gene expression, would be valuable in understanding and addressing a range of prenatally developed diseases and disorders,” Golkaram said.
Generally, he explained, when mRNAs and miRNAs are in the right location and functioning correctly as a “bipartite” network, they inhibit translation of genes and production of proteins to prevent an unhealthy overabundance of any cell type. But if they're not functioning well, such as when a miRNA is suppressed and cannot inhibit gene translation, the result can be overexpression of one or more genes and uncontrolled production of certain cells. Cancer is the most well-known resulting pathology, Golkaram noted.
“In the interaction between these two distinct modes, each messenger RNA can be targeted by multiple micro-RNAs and vice versa,” Golkaram said. “But the interactions of mRNAs and miRNAs do not form a randomly connected network. The network contains communities — you can think of them as clusters — and if you look at a particular cluster that contains a set of both mRNAs and miRNAs and has been extracted from the network, you can see that the cluster is enriched to different extents in different cell types. So, the members of that cluster are the mRNAs and the miRNAs whose expression is known to be regulated up or down in those cell types. If one of your clusters is not working properly, you can tip the balance against or in favor of that cell type, and that can lead to disease.”
Biologists have studied the small, non-coding miRNAs for decades and know that, generally, they serve to suppress excess protein production by preventing mRNA from translating genetic coding. They therefore have a role in fine-tuning the signaling pathways that are related to brain development. But their function depends on context, and it changes as the brain develops and according to where they are expressed.
Until now, no technique has existed for seeing all of the mRNA and miRNA expressions in individual cells. For this project, the researchers used a single-cell qPCR, or quantitative polymerase chain reaction, method. Traditionally, qPCR has been used for populations, not individual cells, Golkaram explained. “You’re looking at a lot of cells at the same time, and the behavior you are capturing is a mean of the population,” he said. “You don't have the resolution to see single cells, which is very important here because we are talking about mRNA-miRNA interactions in specific cell types. So, the mean, or average, behavior of your population is not representative.”
For the project, teams at UC Santa Barbara and UC San Francisco developed a customized qPCR process that could capture both mRNAs and miRNAs in every single cell. “That was an important step forward,” Golkaram said. “It helped us see that the expression levels of these targeting micro-RNAs is highly cell-type specific.”
The scientists used samples from prenatal human brain cells collected under strict ethical and legal regulations. Golkaram, working in the lab of Linda Petzold, his Ph.D. advisor, developed a machine learning approach to identify and extract patterns from what appears to be a chaotic, noisy system containing millions of data points representing the mRNA-miRNA interactions. They then used the patterns to develop hypotheses, which the experimental teams at UCSB and UCSF tested in the lab.
Previous, more general hypotheses have related to the expression of certain types of RNA being higher or lower in specific cell types. “But what we did was to look at all micro-RNAs and mRNAs at the same time, genome-wide, rather than just cherry-picking a few with high precision,” Golkaram explained.
To do that, he said, “We constructed the network by experimentally identifying the set of mRNAs that are micro-RNA targets and connecting all miRNA to their identified targets. That gave us the big picture, allowing us to look at the larger network and the different modules within it. Certain clusters are representative of distinct cell types, and certain micro-RNAs show cell-type-specific expression.
“We can say, ‘OK, in one cell type, the expression of micro RNAs will be modulated in a certain location or at a certain point in time to contribute different levels of regulation in the cluster from which they were extracted,” he continued. “We can therefore compare someone who has autism spectrum disorder with someone who is healthy and see that they have different levels of expression of this miRNA and different levels of regulation.
The researchers used two methods to confirm that a cluster is associated with ASD. One was to compare the mRNA-miRNA levels in that cluster in mice that carry the gene associated with ASD to the levels in healthy mice. The other was to observe significant overlap between the miRNAs and mRNAs in that cluster and those that previous studies of adult humans have shown to be involved in ASD. “We can quantify the difference in the miRNA expression and the mRNA-miRNA interactions in healthy and unhealthy brains,” Golkaram said.
Finally, he added, “Heterogeneity of cell types is critical to an organism’s ability to survive and thrive, and regulating it is the key. If we want to look at different diseases having prenatal origins and understand what causes them, it’s important to have a mechanistic understanding of the mRNA-miRNA interactions. This study tells us that the imbalance between cell types during early development can lead to this disease, and also that this imbalance results from these two modes — mRNA and miRNA — not being correctly aligned.”
The research team at UC Santa Barbara also included Linda R. Petzold, a professor in the departments of computer science and of mechanical engineering; Hongjun R. Zhou (co-first author) and Kylie Huch, both in the Department of Molecular, Cellular and Developmental Biology; and Neha Rani (co-first author), a former postdoctoral fellow in the Kosik lab now at the Indian Institute of Technology in Kanpur, India. Co-first author Tomasz J. Nowakowski is based at UC San Francisco.



