Surviving in Hostile Territory

Many strange creatures live in the deep sea, but few are odder than archaea, primitive single-celled bacteria-like microorganisms. Archaea go to great lengths — eating methane or breathing sulfur or metal instead of oxygen — to thrive in the most extreme environments on the planet.
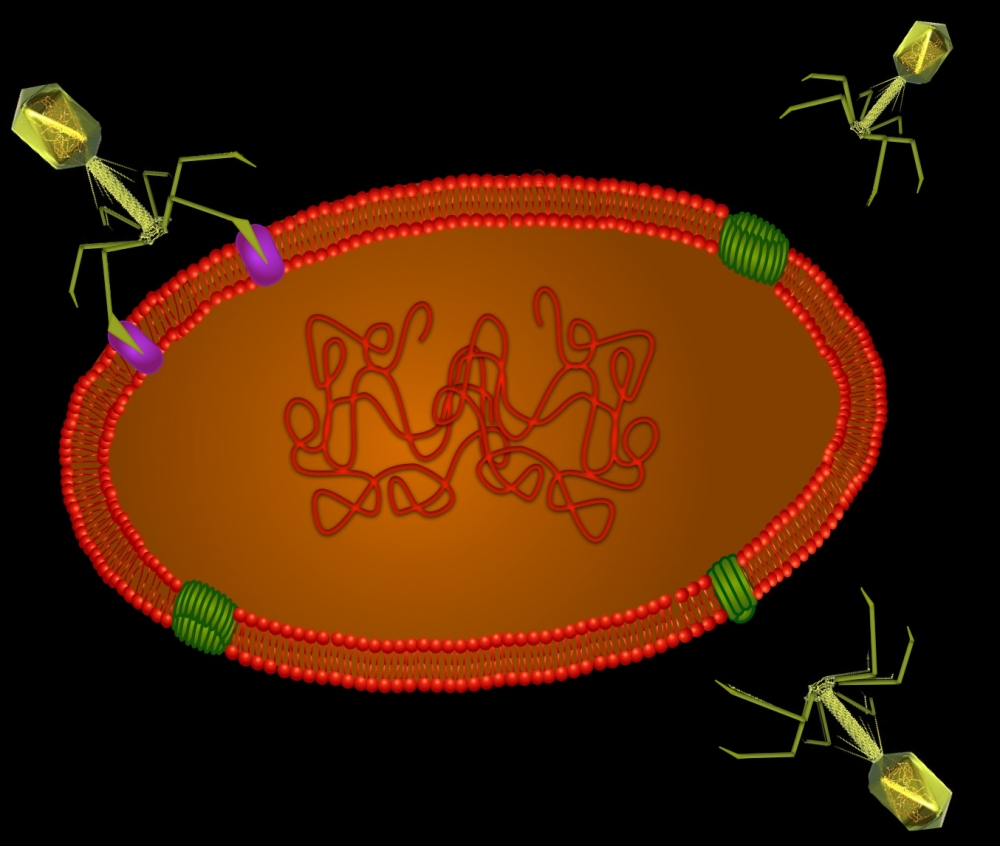
Recently, while searching the ocean’s depths off the coast of Santa Monica, California, a team of UC Santa Barbara scientists discovered something odder still: a remarkable new virus that seemingly infects methane-eating archaea living beneath the ocean’s floor. The investigators were further surprised to discover that this virus selectively targets one of its own genes for mutation and, moreover, that some archaea do too. The researchers’ findings appear today in the journal Nature Communications.
“Our study illustrates that self-guided mutation is relevant to life within the Earth’s subsurface and uncovers mechanisms by which viruses and archaea can adapt in this hostile environment,” said David Valentine, a professor in the Department of Earth Science and at the campus’s Marine Science Institute (MSI). “These findings raise exciting new questions about the evolution and interaction of the microbes that call Earth’s interior home.”
Using the submarine Alvin, Valentine and colleagues collected samples from a deep-ocean methane seep by pushing tubes into the ocean floor and retrieving sediments. The contents were brought back to the lab and fed methane gas, which helped the archaea in the samples grow. When the team assayed the samples for viral infection, they discovered a new virus with a distinctive genetic fingerprint that suggested its likely host was methane-eating archaea.
“It’s now thought that there’s more biomass inside the Earth than anywhere else, just living very, very slowly in this dark, energy-limited, starved environment,” said co-author Sarah Bagby, a postdoctoral scholar in the Valentine lab.
The researchers used the genetic sequence of the new virus to chart other occurrences in global databases. “We found a partial genetic match from methane seeps in Norway and California,” said lead author Blair Paul, a postdoctoral scholar in the Valentine lab. “The evidence suggests this viral type is distributed around the globe in deep ocean methane seeps.”
Further investigation revealed another unexpected finding: a diversity-generating retroelement that greatly accelerates mutation of a specific section of the viral genome. Such small genetic elements had previously been identified in bacteria and their viruses, but never among archaea or the viruses that infect them. While the self-guided mutation element in the archaeal virus clearly resembled the known bacterial elements in many respects, the researchers found that it has a divergent evolutionary history.
“The target of guided mutation — the tips of the virus that make first contact when infecting a cell — was similar,” said Paul. “The ability to mutate those tips is an offensive countermeasure against the cell’s defenses — a move that resembles a molecular arms race.”
Having found guided mutation in a virus infecting archaea, the scientists reasoned that archaea themselves might use the same mechanism for genetic adaptation. Indeed, in an exhaustive search, they identified parallel features in the genomes of a more mysterious subterranean group of archaea known as nanoarchaea. Unlike the deep-ocean virus that uses guided mutation to alter a single gene, nanoarchaea target at least four distinct genes.
“This is a new record,” said Bagby. “Previously, a few bacteria had been observed to target two genes with this mechanism. That may not seem like a huge difference, but targeting four is extraordinary. If they’re all firing at once, suddenly the number of combinations of protein variants in play is really massive.”
According to Valentine, the genetic mutation that engenders these potential variations may be a key element to survival of archaea beneath the Earth’s surface. “The cell is choosing to modify certain proteins,” he explained.“It’s doing its own protein engineering internally. While we don’t know what those proteins are being used for, I think learning about the process can tell us something about the environment in which these organisms thrive. Right now, we know so little about life in that environment.”
This research was supported by an NSF Dimensions of Biodiversity grant aimed at characterizing virus-mediated microbial diversity in methane seep ecosystems. Viral DNA sequencing was provided through a Gordon and Betty Moore Foundation grant to the Broad Institute. The research team also included scientists from UCLA, UC San Diego and the Department of Energy’s Joint Genome Institute.