In Protein Folding, Internal Friction May Play a More Significant Role than Previously Thought


An international team of researchers has reported a new understanding of a little-known process that happens in virtually every cell of our bodies.
Protein folding is the process by which not-yet folded chains of amino acids assume their specific shapes, hence taking on their specific functions. These functions vary widely: In the human body, proteins fold to become muscles, hormones, enzymes, and various other components.
"This protein folding process is still a big mystery," said UC Santa Barbara physicist Everett Lipman, one of several authors of a paper, "Quantifying internal friction in unfolded and intrinsically disordered proteins with single-molecule spectroscopy." The paper was published in the Proceedings of the National Academy of Sciences.
A protein's final shape, said Lipman, is primarily determined by the sequence of amino acid components in the unfolded chain. In the process, the components bump up against each other, and when the right configuration is achieved, the chain passes through its "transition state" and snaps into place.
"What we would like to understand eventually is how the chemical sequence of a protein determines what it is going to become and how fast it is going to get there," Lipman said.
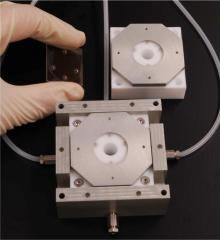
Using a microfluidic mixing technique pioneered in the UCSB physics department by former graduate student Shawn Pfeil, the research team, including collaborators from the University of Zurich and the University of Texas, was able to monitor extremely rapid reconfiguration of individual protein molecules as they folded.
In the microfluidic mixer, a "denaturant" chemical used to unravel the proteins was quickly diluted, enabling observation of folding under previously inaccessible natural conditions. The measurements demonstrated that internal friction plays a more significant role in the folding process than could be seen in prior experiments, especially when the protein starts in the more compact unfolded configuration it would have in a denaturant-free living cell.
"At those size scales, everything is dominated by friction," said Lipman, comparing the environment of a protein molecule in water to a human body in molasses. Friction between the molecule and its liquid environment is an issue, as well as the "dry" friction that is independent of the surrounding solvent.
Internal friction slows down the folding process by reducing the rate at which the amino acid chain explores different configurations that may lead to the transition state.
The longer it takes to find its native state –– its final form –– the higher the likelihood it could get stuck in an unfolded state.
"When it is unfolded, it is more vulnerable to being trapped in a misfolded state, or to aggregation with other unfolded protein molecules," said Lipman. Aggregation of misfolded proteins is thought to be a contributor to many types of diseases, such as the amyloid plaques that are associated with Alzheimer's disease. Alternatively, the unfolded and not usable protein could be broken back up into its component amino acids by the cell.
While there is no confirmed link between internal friction and aggregation, or any pattern of friction for one protein that affects others in the same way, Lipman and his colleagues are getting closer to understanding the degree to which internal friction affects the protein folding process.
"These measurements show that under realistic conditions, internal friction plays a significant role in the dynamics of the unfolded state. If a model of the protein folding process doesn't account for this, it will need to be reconsidered," he said.
Related Links



