Why Antibiotics Fail

When a patient is prescribed the wrong antibiotic to treat a bacterial infection, it’s not necessarily the physician who is at fault. The current antibiotic assay — standardized in 1961 by the World Health Organization and used worldwide — is potentially flawed.
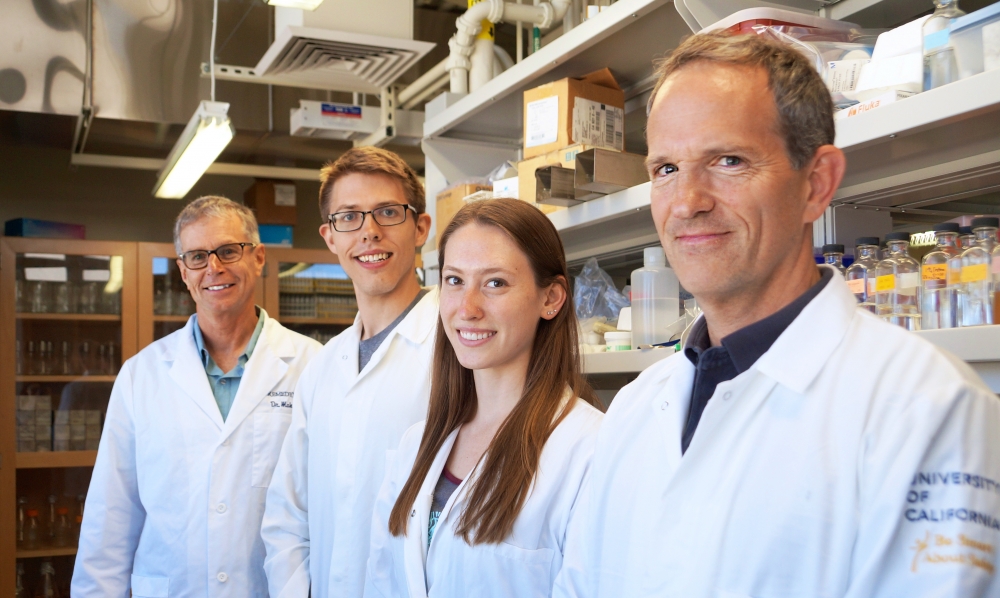
So says UC Santa Barbara biologist Michael Mahan, whose lab has developed a new antimicrobial susceptibility test that could transform the way antibiotics are developed, tested and prescribed.
The standard test specifies how well drugs kill bacteria on petri plates containing Mueller-Hinton Broth, a nutrient-rich laboratory medium that fails to reproduce most aspects of a natural infection. Now, Mahan and colleagues have used a mouse model to demonstrate that a variety of antibiotics work differently against various pathogens when inside the mammalian body. Their findings appear in the journal EBioMedicine.
“The message is simple: Physicians may be relying on the wrong test for identifying antibiotics to treat infections,” said Mahan, a professor in the Department of Molecular, Cellular and Developmental Biology. “By developing a test that mimics conditions in the body, we have identified antibiotics that effectively treat infections caused by diverse bacteria, including MRSA, the cause of deadly Staphylococcal infections. These drugs have been overlooked because they failed the standard tests, despite being inexpensive, nontoxic and available at local pharmacies.”
The research has significant implications for public health. If a drug that passed the standard test doesn’t work, physicians can now choose a different drug immediately rather than increase the dose of the same drug when patients return — often in worse condition — after an ineffective first course of treatment.
Reliance on the standard test may have contributed to the rise in multidrug-resistant bacteria, Mahan noted, due to the continued prescription of ineffective antibiotics. Further, he added, the standard test may also be slowing the discovery of new antibiotics. “These ‘wonder drugs’ may already exist but have been rejected by the standard test and are consequently not used in practice,” Mahan said.
The scientists also report a way to “fix” the standard test to better predict how well antibiotics will treat infections: Simply add sodium bicarbonate. More commonly known as baking soda, this chemical is found in abundance in the body, where it helps to maintain precise blood pH. “Sodium bicarbonate makes the petri plates behave more like the body and increases the test’s accuracy for assigning the appropriate antibiotic to treat infections,” explained co-lead author Douglas Heithoff, a project scientist at the Center for Nanomedicine.
Mahan also points out that pharmaceutical companies could benefit from using the revised test to rescreen their collections of purified compounds that have failed the standard test. “There could be a treasure trove of compounds that have been shelved but could actually be quite effective against antibiotic-resistant strains,” he said.
“Things aren’t as gloomy as we thought,” Mahan added. “We just have to be smart about it and change the way we’re using the drugs we already have while we continue to search for new ones.”



